Running interactive Optuna studies
In this tutorial, we deploy a function service that hosts an interactive Optuna study. This lets us run trials on Covalent Cloud compute resources while accessing the study and its outcomes in real-time through a custom API.
To prepare a local environment for this tutorial, run the following command.
pip install "covalent-cloud>=0.84.1" "optuna==4.0.0" "pandas==2.2.3"
Imports
Let's start with a cell to import the necessary libraries.
import base64
import json
import pickle
import time
from pathlib import Path
from pprint import pprint
import covalent as ct
import covalent_cloud as cc
import optuna
import pandas as pd
from requests.exceptions import HTTPError
CC_API_KEY = cc.get_api_key()
TIME_LIMIT = "7 days"
Target Function
For the purposes of this tutorial, we use a simple target function as a stand-in for the complicated loss surface. Applying Covalent Cloud decorators to this function then makes it a cloud executable workflow.
For a more practical optimization, the environment and executor here should be adjusted accordingly.
TRIAL_ENV = 'optuna-trial-env-1'
# no extra dependencies needed for this example
cc.create_env(name=TRIAL_ENV, pip=[], wait=True)
trial_executor = cc.CloudExecutor(env=TRIAL_ENV, num_cpus=2, memory='2GB')
# cloud executable one-task workflow.
@ct.lattice(executor=trial_executor, workflow_executor=trial_executor)
@ct.electron
def target_func(x, y):
return 0.5 * x**2 + (0.5 * y**2 - 1)**3 + 1
Notice that this target function has two unstable local minima at (0, +sqrt(2)) and (0, -sqrt(2)), and one stable global minimum at (0, 0). This is not something we'd know a priori in a real-world optimization problem, of course, but we'll exploit this knowledge for the sake of this tutorial.

Optuna study service
Utilities
Here we have a helper class and some utility functions that streamline our service definition. It is not necessary to do things this way, but sometimes it helps with neatness.
The code cell below introduces the following functionality:
- a helper class to track Covalent workflows corresponding to study trials.
- some helper functions to allow JSON-serializable communication of important data.
With Covalent Cloud, we can define things in the global scope and simply refer to them in our service code. All the necessary components are automatically serialized and communicated to remote jobs.
class StudyHelper:
def __init__(self):
self.root_id = None
self.submitted = {}
self.completed = {}
self.failed = {}
def tell(self, study):
"""Check statuses, tell complete studies, return status."""
newly_completed = []
newly_failed = []
running = []
for _id, trial in self.submitted.items():
if _id in self.completed or _id in self.failed:
continue
try:
# Get covalent workflow result.
r = cc.get_result(_id).status.STATUS.value
except HTTPError:
continue
# Update the status for response.
if r == "COMPLETED":
self.completed[_id] = trial
newly_completed.append(_id)
elif r == "FAILED":
self.failed[_id] = trial
newly_failed.append(_id)
elif r == "RUNNING":
running.append([trial.number, _id])
# Tell study about completed trials.
for _id in newly_completed:
trial = self.submitted.pop(_id)
result = cc.get_result(_id, wait=True).result.load()
study.tell(trial, result)
# Tell study about failed trials.
for _id in newly_failed:
trial = self.submitted.pop(_id)
study.tell(trial, None)
return self.content(running=running)
def content(self, **extras):
"""Return the content of the helper."""
return {
"root_id": self.root_id,
"submitted": {t.number: _id for _id, t in self.submitted.items()},
"completed": {t.number: _id for _id, t in self.completed.items()},
"failed": {t.number: _id for _id, t in self.failed.items()}, **extras,
}
def pickle_study(study_obj):
"""Pickle a study and make serializable."""
study_bytes = pickle.dumps(study_obj)
return base64.b64encode(study_bytes).decode()
def unpickle_study(study_bytes_str):
"""Unpickle a serializable study."""
study_bytes = base64.b64decode(study_bytes_str)
return pickle.loads(study_bytes)
def get_distributions(dists):
distributions = {}
for k, v in dists.items():
cls = getattr(optuna.distributions, v[0])
distributions[k] = cls(*v[1])
return distributions
Definition
We first create an environment and a suitable executor. The service itself requires little compute, so a couple CPUs will more than suffice.
SERVICE_ENV = "optuna-service"
cc.create_env(
name=SERVICE_ENV,
pip=["numpy==1.23.5", "optuna==4.0.0", "pandas==2.2.3"],
wait=True,
)
service_ex = cc.CloudExecutor(
env=SERVICE_ENV,
num_cpus=2,
memory="10GB",
time_limit=TIME_LIMIT,
)
The service itself is defined as follows, including arbitrary endpoints to create our desired API.
The initializer function (i.e. the optuna_interface function, decorated with @cc.service) creates a new Optuna study and a helper object to track the status of trials. Every item in the returned dictionary is accessible to any service endpoint that declares an argument with a matching name.
@cc.service(executor=service_ex, name="Optuna Interface")
def optuna_interface(trial_lattice, **create_study_kwargs):
cc.save_api_key(CC_API_KEY)
return {
"study": optuna.create_study(**create_study_kwargs),
"helper": StudyHelper(),
"trial_lattice": trial_lattice,
} # auto-passed to endpoints below, as required
Once decorated with @cc.service, the initializer itself becomes a decorator with which we can define the service's endpoints.
@optuna_interface.endpoint("/ask")
def ask_and_run(study, helper, trial_lattice, *, dists) -> str:
"""Create a new trial and run it."""
fixed_distributions = get_distributions(dists)
trial = study.ask(fixed_distributions=fixed_distributions)
if helper.root_id:
_id = cc.redispatch(helper.root_id)(**trial.params)
else:
_id = cc.dispatch(trial_lattice)(**trial.params)
helper.root_id = _id
helper.submitted[_id] = trial
return _id
@optuna_interface.endpoint("/tell")
def tell(study, helper) -> dict:
"""
Check statuses, tell complete studies, return status.
"""
return helper.tell(study)
@optuna_interface.endpoint("/trials")
def get_trials(study) -> dict:
"""
Get a dictionary of current trials.
"""
return json.loads(study.trials_dataframe().to_json())
@optuna_interface.endpoint("/best")
def get_best(study: optuna.Study, keys=None) -> dict:
"""
Get the best value and parameters so far.
"""
return {"value": study.best_value, "params": study.best_params}
@optuna_interface.endpoint("/all-done")
def check_all_done(study, helper) -> bool:
"""
Return 1 if all trials are done, 0 otherwise.
"""
status = helper.tell(study)
return int(len(status["submitted"]) == len(status["running"]) == 0)
@optuna_interface.endpoint("/content")
def get_content(helper) -> dict:
"""
Get the content of the helper.
"""
return helper.content()
@optuna_interface.endpoint("/study")
def get_study(study) -> str:
"""
Get the study object as a serialized byte string.
"""
return pickle_study(study) # `unpickle_study` to recover object client-side
Deploy the service
To start the service, we deploy it in the usual way.
# Deploy with custom initializer arguments.
client = cc.deploy(optuna_interface)(
trial_lattice=target_func,
direction="minimize",
study_name="test-study"
)
# Wait for service to reach an active state.
client = cc.get_deployment(client, wait=True)
print(client)
╭──────────────────────────────── Deployment Information ────────────────────────────────╮
│ Name Optuna Interface │
│ Description Add a docstring to your service function to populate this section. │
│ Function ID 67409941e529fcff11cdadb0 │
│ Address https://fn-a.prod.covalent.xyz/67409941e529fcff11cdadb0 │
│ Status ACTIVE │
│ Auth Enabled Yes │
╰────────────────────────────────────────────────────────────────────────────────────────╯
╭────────────────────────────────────── Endpoints ───────────────────────────────────────╮
│ Route POST /ask │
│ Streaming No │
│ Description Create a new trial and run it. │
│ │
│ Route POST /tell │
│ Streaming No │
│ Description │
│ Check statuses, tell complete studies, return status. │
│ │
│ │
│ Route POST /trials │
│ Streaming No │
│ Description │
│ Get a dictionary of current trials. │
│ │
│ │
│ Route POST /best │
│ Streaming No │
│ Description │
│ Get the best value and parameters so far. │
│ │
│ │
│ Route POST /all-done │
│ Streaming No │
│ Description │
│ Return 1 if all trials are done, 0 otherwise. │
│ │
│ │
│ Route POST /content │
│ Streaming No │
│ Description │
│ Get the content of the helper. │
│ │
│ │
│ Route POST /study │
│ Streaming No │
│ Description │
│ Get the study object as a serialized byte string. │
│ │
╰────────────────────────────────────────────────────────────────────────────────────────╯
Authorization token: <authorization-token-redacted>
Using the service
Running an initial set of trials
Let's use the /ask endpoint to run a few initial trials. We'll run 8 trials in this example, using a distribution of x and y values near one of the local minima.
d1 = {
"x": ("FloatDistribution", (-2.0, 2.0)),
"y": ("FloatDistribution", (-3.0, -1.0)),
}
# Run 8 trials with sampled parameters.
asked = []
for _ in range(8):
asked.append(client.ask(dists=d1))
time.sleep(1)
asked
['86b6dd82-274c-4962-b56d-a4c46bfbb694'
'cce858df-51e1-4b23-b0f2-c846b4518142'
'3f7f4c54-dc3e-46e3-8c41-a404f73cd4a5'
'bb2b93de-6c53-4d5b-a333-92dfa98602e2'
'6667fd38-8678-4399-b02d-451470ddee8a'
'fadb8910-46f0-4916-83d2-df98c9929b94'
'4c2ac385-bd5a-4ee1-bffb-be4ddab392fc'
'1b69098f-5b30-4b77-9026-df229e802e76']
Checking the status of initial trials
The /tell endpoint (logic defined inside StudyHelper) will check the status of all trials and return the completed ones. We can use this to check the status of our initial trials.
Depending on the amount of time that has passed, the output may look something like this:
pprint(client.tell())
{'completed': {'0': '86b6dd82-274c-4962-b56d-a4c46bfbb694'},
'failed': {},
'root_id': '86b6dd82-274c-4962-b56d-a4c46bfbb694',
'running': {'1': 'cce858df-51e1-4b23-b0f2-c846b4518142',
'2': '3f7f4c54-dc3e-46e3-8c41-a404f73cd4a5',
'3': 'bb2b93de-6c53-4d5b-a333-92dfa98602e2'},
'submitted': {'4': '6667fd38-8678-4399-b02d-451470ddee8a',
'5': 'fadb8910-46f0-4916-83d2-df98c9929b94',
'6': '4c2ac385-bd5a-4ee1-bffb-be4ddab392fc',
'7': '1b69098f-5b30-4b77-9026-df229e802e76'}}
We can also use the /all-done endpoint as follows, to wait until all trials are done (or submit additional trials in the meantime).
Waiting for all trials to finish
# wait for all trials to finish
finished = client.all_done()
while not finished:
time.sleep(5)
finished = client.all_done()
Using the /trials endpoint, we can obtain a summary of the trials that have been run so far. The response here can be converted directly into a pandas DataFrame for easier inspection.
Recovering a DataFrame of trials
The /trials endpoint returns a dictionary of the study.trials_dataframe() method's output. We can convert this back into a DataFrame for easier inspection.
pd.DataFrame.from_dict(client.trials())
| number | value | datetime_start | datetime_complete | duration | params_x | params_y | state |
|---|---|---|---|---|---|---|---|
| 0 | 1.609996 | 1732286939055 | 1732287584351 | 645296 | 1.11195 | -1.263473 | COMPLETE |
| 1 | 4.866197 | 1732286941294 | 1732287584633 | 643339 | 0.354487 | -2.263162 | COMPLETE |
| 2 | 1.28533 | 1732286944124 | 1732287584942 | 640817 | 0.722207 | -1.606607 | COMPLETE |
| 3 | 2.727394 | 1732286945678 | 1732287585218 | 639539 | -1.884925 | -1.125945 | COMPLETE |
| 4 | 1.566873 | 1732286949468 | 1732287585448 | 635979 | 1.064877 | -1.380043 | COMPLETE |
| 5 | 37.481463 | 1732286951763 | 1732287585743 | 633979 | 1.047317 | -2.932534 | COMPLETE |
| 6 | 17.990337 | 1732286953219 | 1732287586033 | 632814 | -0.953944 | -2.663696 | COMPLETE |
| 7 | 15.377415 | 1732286954797 | 1732287586321 | 631524 | 1.296379 | -2.601262 | COMPLETE |
Checking best value and parameters so far
At this point, /best endpoint on our service can give us the best parameters and value found so far.
client.best(keys=["params", "value"])
{'value': 1.285330345491837,
'params': {'x': 0.7222071617109473, 'y': -1.6066065739414577}}
Considering our input distributions for these initial trials, we are indeed converging upon the local minimum value at (0, -sqrt(2)).
Running additional trials
Let's try searching for the global minimum, which we happen to know is at (0, 0). We'll run 5 more trials with a distribution centered around this point.
d2 = {
"x": ("FloatDistribution", (-.5, .5)),
"y": ("FloatDistribution", (-.5, .5)),
}
# Run 5 more trials with sampled parameters.
asked = []
for _ in range(5):
asked.append(client.ask(dists=d2))
time.sleep(1)
asked
['9d2ec6f5-750c-4086-af13-746df722fbf7',
'd4b45e38-4e1d-4f55-af20-4c52acf29f93',
'78ed0926-2b19-4ef9-839c-5f61c88174a8',
'9d78726c-42f1-4b0c-b593-c3767b1be089',
'f817e9e7-e881-487a-87cc-f7010cae8782']
Checking the status of additional trials
Shortly after, we can see via the /tell endpoint that additional trials are under way.
pprint(client.tell())
{'completed': {'0': '86b6dd82-274c-4962-b56d-a4c46bfbb694',
'1': 'cce858df-51e1-4b23-b0f2-c846b4518142',
'2': '3f7f4c54-dc3e-46e3-8c41-a404f73cd4a5',
'3': 'bb2b93de-6c53-4d5b-a333-92dfa98602e2',
'4': '6667fd38-8678-4399-b02d-451470ddee8a',
'5': 'fadb8910-46f0-4916-83d2-df98c9929b94',
'6': '4c2ac385-bd5a-4ee1-bffb-be4ddab392fc',
'7': '1b69098f-5b30-4b77-9026-df229e802e76'},
'failed': {},
'root_id': '86b6dd82-274c-4962-b56d-a4c46bfbb694',
'running': [[8, '9d2ec6f5-750c-4086-af13-746df722fbf7'],
[9, 'd4b45e38-4e1d-4f55-af20-4c52acf29f93'],
[10, '78ed0926-2b19-4ef9-839c-5f61c88174a8']],
'submitted': {'10': '78ed0926-2b19-4ef9-839c-5f61c88174a8',
'11': '9d78726c-42f1-4b0c-b593-c3767b1be089',
'12': 'f817e9e7-e881-487a-87cc-f7010cae8782',
'8': '9d2ec6f5-750c-4086-af13-746df722fbf7',
'9': 'd4b45e38-4e1d-4f55-af20-4c52acf29f93'}}
Waiting for all trials to finish
We can use the /all-done endpoint once again to wait for all new trials to finish.
# wait for all trials to finish
finished = client.all_done()
while not finished:
time.sleep(5)
finished = client.all_done()
Checking best value and parameters overall
The best outcome so far will now include all the studies so far. We can see here that the best parameters a now closer to the global minimum at (0, 0).
client.best(keys=["params", "value"])
{'value': 0.1170285789683333,
'params': {'x': 0.4294500956067753, 'y': 0.12915887785342173}}
Recovering the study object
Thanks to the /study endpoint, we can recover the study Python object and inspect it further. This lets us access non-JSON-serializable data like the study's actual trial objects etc.
study_str = client.study()
study = unpickle_study(study_str)
print(study.best_trials)
FrozenTrial(number=0, state=TrialState.COMPLETE, values=[1.609996005517527], datetime_start=datetime.datetime(2024, 11, 22, 14, 48, 59, 55562), datetime_complete=datetime.datetime(2024, 11, 22, 14, 59, 44, 351807), params={'x': 1.1119498168408013, 'y': -1.2634727129721934}, user_attrs={}, system_attrs={}, intermediate_values={}, distributions={'x': FloatDistribution(high=2.0, log=False, low=-2.0, step=None), 'y': FloatDistribution(high=-1.0, log=False, low=-3.0, step=None)}, trial_id=0, value=None)
FrozenTrial(number=1, state=TrialState.COMPLETE, values=[4.866197352815817], datetime_start=datetime.datetime(2024, 11, 22, 14, 49, 1, 294100), datetime_complete=datetime.datetime(2024, 11, 22, 14, 59, 44, 633116), params={'x': 0.3544874553422792, 'y': -2.2631621517141465}, user_attrs={}, system_attrs={}, intermediate_values={}, distributions={'x': FloatDistribution(high=2.0, log=False, low=-2.0, step=None), 'y': FloatDistribution(high=-1.0, log=False, low=-3.0, step=None)}, trial_id=1, value=None)
FrozenTrial(number=2, state=TrialState.COMPLETE, values=[1.285330345491837], datetime_start=datetime.datetime(2024, 11, 22, 14, 49, 4, 124876), datetime_complete=datetime.datetime(2024, 11, 22, 14, 59, 44, 942268), params={'x': 0.7222071617109473, 'y': -1.6066065739414577}, user_attrs={}, system_attrs={}, intermediate_values={}, distributions={'x': FloatDistribution(high=2.0, log=False, low=-2.0, step=None), 'y': FloatDistribution(high=-1.0, log=False, low=-3.0, step=None)}, trial_id=2, value=None)
...
Covalent UI
Since trials correspond to Covalent workflows, we can inspect the status of each computation in the Covalent UI. Our service tracks the root workflow ID that identifies all the above experiments:
client.content()["root_id"]
'86b6dd82-274c-4962-b56d-a4c46bfbb694'
This unique ID together with the workflow name (target_func) helps us locate the root workflow in the Covalent UI.
Redispatch
You may have noticed that we used cc.redispatch inside our /ask endpoint after the very first trial was run with cc.dispatch. This helps us organize related dispatches under a single root workflow, making it easier to track and manage them.
To access the table of re-dispatched workflows, simply click on the "+12" icon in the root workflow's row.
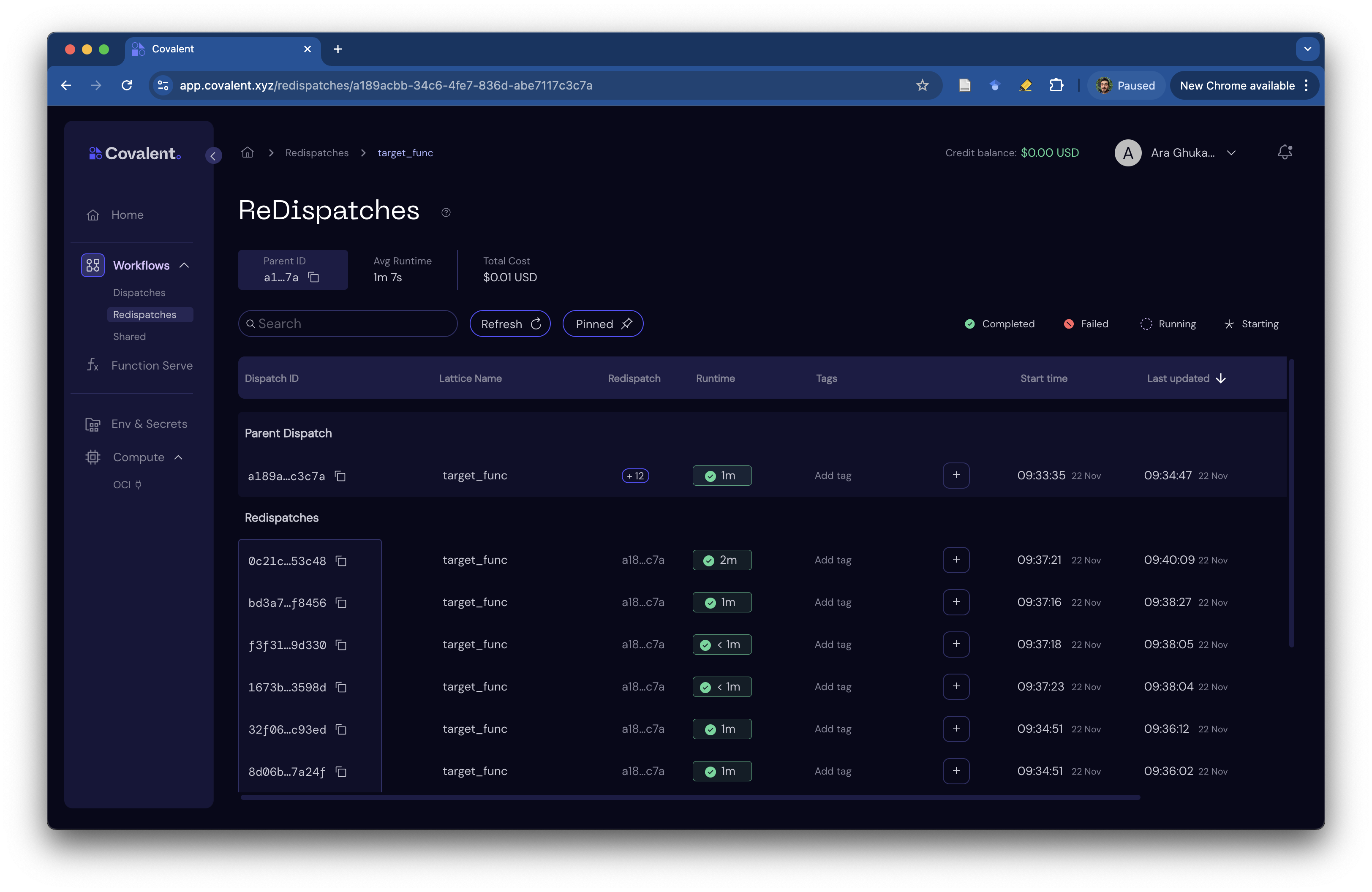
Clean up
The service can be torn down any time before its executor's time limit by running the following.
client.teardown()