Agentic Retrieval-Augmented Generation with Snowflake and Llama3
Retrieval-Augmented Generation (RAG) combines the capabilities of large language models with a retrieval mechanism for incorporating relevant information from external documents or databases. This approach can improve the accuracy and relevance of responses, especially for queries requiring up-to-date or domain-specific knowledge.
In this tutorial, we'll build a question answering RAG application using Llama3 and a Snowflake database.
Outline
Infrastructure setup
Agentic RAG application
- Building the RAG application
- Evaluating a single example
- Evaluating many examples
- Cleaning up
- Conclusion
Setup
import json
import os
import shutil
import subprocess
from pathlib import Path
from typing import Any, Dict, List, Optional, Union
import covalent as ct
import covalent_cloud as cc
from covalent_cloud.function_serve.deployment import Deployment
# cc.save_api_key("<your-covalent-cloud-api-key>")
To start, let's store our Snowflake credentials as secrets in our Covalent Cloud account. We can also save a Hugging Face token in order to access gated repositories.
Each secret below will be available as an environment variable inside all running tasks.
cc.store_secret(name="SNOWFLAKE_USER", value="<your-snowflake-username>")
cc.store_secret(name="SNOWFLAKE_PASSWORD", value="<your-snowflake-password>")
cc.store_secret(name="SNOWFLAKE_ACCOUNT", value="<your-snowflake-account>")
cc.store_secret(name="HF_TOKEN", value="<your-hf-token>")
We'll use the following cloud environment for this tutorial.
ENV_NAME = "AgentRAG"
cc.create_env(
name=ENV_NAME,
pip=[
"covalent-cloud==0.72.0rc0",
"datasets==2.20.0",
"lm-format-enforcer==0.10.4",
"snowflake==0.10.0",
"torch==2.3.0",
"tqdm==4.66.4",
"vllm==0.5.1",
],
wait=True
)
Additionally, the cloud volume below will serve as persistent storage for our model and dataset files.
cloud_volume = cc.volume("agent-rag")
Compute resources
Without too much granularity, we'll utilize the following two resource specifications for our tasks and services. In Covalent, these resources are represented by Cloud Executors.
Below we instantiate a lightweight CPU-only executor and a GPU-enabled executor that specifies one NVIDIA A100. The latter specification (i.e. the gpu_A100x1 executor) will be useful for tasks that require an LLM.
cpu_x2 = cc.CloudExecutor(
env=ENV_NAME,
num_cpus=2,
memory="18GB",
time_limit="2 hours",
)
gpu_A100x1 = cc.CloudExecutor(
env=ENV_NAME,
num_cpus=2,
num_gpus=1,
gpu_type="a100-80g",
memory="48GB",
time_limit="6 hours",
)
Snowflake connection helper
This is a normal Python function. Because it exists in the global context, our Covalent building blocks (i.e. functions with @ct.electron or @cc.service on top) will have access to it when they execute remotely.
This function connects to Snowflake and initializes a table for our vector embeddings.
def _connect_to_snowflake(
user: str,
password: str,
account: str,
warehouse_name: str,
database_name: str,
schema_name: str,
table_name: str,
embed_size: int,
) -> Any:
"""Initialize and connect to a Snowflake DB."""
import snowflake.connector
conn = snowflake.connector.connect(
user=user,
password=password,
account=account
)
# Set up the Snowflake DB.
cursor = conn.cursor()
cursor.execute(f"CREATE WAREHOUSE IF NOT EXISTS {warehouse_name}")
cursor.execute(f"CREATE DATABASE IF NOT EXISTS {database_name}")
cursor.execute(f"USE DATABASE {database_name}")
cursor.execute(f"CREATE SCHEMA IF NOT EXISTS {schema_name}")
cursor.execute(f"USE SCHEMA {database_name}.{schema_name}")
# Create the table if it does not exist.
cursor.execute(
f"CREATE TABLE IF NOT EXISTS {table_name} "
f"(text STRING, embedding VECTOR(float, {embed_size}))"
)
return conn
Covalent tasks
Downloading models and data
Here's our first Covalent task. It downloads Hugging Face repos containing datasets or models and stores them in our cloud volume. The cpu_2x executor is sufficient here, since none of this requires GPUs.
@ct.electron(executor=cpu_x2)
def download_hf_repo(repo_id: str, repo_type: str, overwrite: bool) -> str:
"""Download a Hugging Face repository and save it to cloud storage.
Args:
repo_id: Repository ID.
repo_type: Repository type, either 'model' or 'dataset'.
overwrite: Whether to overwrite existing stored repo files.
Returns:
Path of the downloaded repo in cloud storage.
"""
save_basedir = Path("/tmp")
save_name = repo_id.replace("/", "__")
local_path = save_basedir / save_name # local path
cloud_path = cloud_volume / save_name # cloud storage path
hf_cmd = (
"huggingface-cli download {repo_id} "
"--local-dir {save_path} "
f"--repo-type {repo_type} "
"--quiet"
)
if overwrite or not cloud_path.exists():
# Download the repo using Hugging Face CLI.
subprocess.run(
hf_cmd.format(repo_id=repo_id, save_path=local_path),
shell=True,
check=True,
)
# Save to cloud storage.
print(f"Writing files to {cloud_path}")
shutil.copytree(local_path, cloud_path, dirs_exist_ok=True)
return str(cloud_path)
Populating embeddings in the vector DB
This next Covalent task populates our Snowflake table with vector embeddings for our dataset. Computing the embeddings requires an embedding model, and therefore a GPU. To accommodate this, we use the gpu_A100x1 executor for this task.
@ct.electron(executor=gpu_A100x1)
def populate_db(
embedding_model_path: str,
dataset_path: str,
dataset_name: str,
dataset_split: str,
dataset_embed_feature: str,
warehouse_name: str,
database_name: str,
schema_name: str,
table_name: str,
embed_size: int,
min_items: int = 1000,
) -> None:
"""Populate a Snowflake database with embeddings from a text dataset.
Args:
embedding_model_path: Cloud path to the embedding model files.
dataset_path: Cloud path to the dataset files.
dataset_name: Name of the dataset.
dataset_split: Dataset split string, e.g. 'train[:40%]'.
dataset_embed_feature: Name of the feature to embed in the dataset.
warehouse_name: Snowflake warehouse name.
database_name: Snowflake database name.
schema_name: Snowflake schema name.
table_name: Snowflake table name.
embed_size: Size of the embedding vectors.
min_items: Skip populating the database if at least this many items exist. Defaults to 1000.
Returns:
A dictionary containing the Snowflake connection parameters.
"""
from datasets import load_dataset_builder
from tqdm import tqdm
from vllm import LLM
# Load the embedding model.
embedding_model = LLM(embedding_model_path)
# Load the dataset.
if dataset_name:
builder = load_dataset_builder(dataset_path, dataset_name)
else:
builder = load_dataset_builder(dataset_path)
builder.download_and_prepare()
data = builder.as_dataset(split=dataset_split)
# Extract the text field.
data_text = data[dataset_embed_feature]
# Encode the entire text dataset.
outputs = embedding_model.encode(data_text)
# Connect to Snowflake.
credentials = {
# Account secrets are available as env vars in running tasks.
"user": os.environ["SNOWFLAKE_USER"],
"password": os.environ["SNOWFLAKE_PASSWORD"],
"account": os.environ["SNOWFLAKE_ACCOUNT"],
}
connect_params = {
"warehouse_name": warehouse_name,
"database_name": database_name,
"schema_name": schema_name,
"table_name": table_name,
"embed_size": embed_size,
}
db_conn = _connect_to_snowflake(**{**credentials, **connect_params})
cursor = db_conn.cursor()
# Check if table is already populated.
count = cursor.execute(f"SELECT COUNT(*) FROM {table_name}").fetchone()[0]
if count < min_items:
insert_cmd = (
f"INSERT INTO {table_name}(text, embedding) "
f"SELECT '{{text}}', {{embedding}}::VECTOR(FLOAT, {embed_size})"
)
for text, output in tqdm(zip(data, outputs)):
embedding = output.outputs.embedding
escaped_text = json.dumps(text)
try:
cursor.execute(
insert_cmd.format(text=escaped_text, embedding=embedding)
)
except Exception as e:
print(f"Error inserting '{text}': {e}")
return connect_params
Covalent services
Services in Covalent, unlike tasks, are long-running and interactive processes. Services integrate with workflows in the same way as tasks. They are useful for hosting models or anything else that's slow to initialize and/or interacted with multiple times.
We'll employ two services in this tutorial, one to host our embedding model and another to host our generation model.
Embedding service
This service hosts the embedding model, which is used to compute vector embeddings for unseen text inside retrieval queries. Once we obtain the embedding vector for this text, database documents are selected based on the cosine similarity between their vector and the text's vector.
@cc.service(executor=gpu_A100x1, name="Embedding Service", volume=cloud_volume)
def embedding_service(
model_path: str,
connect_params: Dict[str, Any],
):
"""Initialize the embedding service and embed a dataset as
text-vector pairs in a Snowflake DB.
Args:
model_path: Storage path of the embedding model.
connect_params: Parameters to connect to the Snowflake DB.
"""
from vllm import LLM
# Load the model.
embedding_model = LLM(model_path)
# Connect to Snowflake.
credentials = {
# Account secrets are available as env vars for running tasks.
"user": os.environ["SNOWFLAKE_USER"],
"password": os.environ["SNOWFLAKE_PASSWORD"],
"account": os.environ["SNOWFLAKE_ACCOUNT"],
}
db_conn = _connect_to_snowflake(**{**credentials, **connect_params})
embed_size = connect_params["embed_size"]
table_name = connect_params["table_name"]
# Partially formatted SQL command to retrieve the most similar text.
retrieve_cmd = (
"SELECT text, "
f"VECTOR_COSINE_SIMILARITY(embedding, {{target}}::VECTOR(FLOAT, {embed_size})) "
f"AS similarity FROM {table_name} "
"ORDER BY similarity DESC "
"LIMIT {limit}"
)
return {
"embedding_model": embedding_model,
"db_conn": db_conn,
"retrieve_cmd": retrieve_cmd
}
This service's /retrieve endpoint, defined below, is called to retrieve documents most similar to the given text.
@embedding_service.endpoint("/retrieve")
def retrieve(
embedding_model=None,
db_conn=None,
retrieve_cmd=None,
*,
text=None,
num_results=3,
) -> str:
"""Retrieve the most similar text from the Snowflake DB.
Kwargs:
text: Input text to find the most similar text.
num_results: Number of results to return. Defaults to 3.
Returns:
The most similar text from the Snowflake DB.
"""
if text is None:
return "Please provide a text input."
# Encode the input text.
output = embedding_model.encode([text])
embedding = output[0].outputs.embedding
# Retrieve the most similar text from the Snowflake DB.
cursor = db_conn.cursor()
cursor.execute(
retrieve_cmd.format(target=embedding, limit=num_results)
)
return [t[0].strip("\"'") for t in cursor.fetchall()]
Generator service
The generator service hosts a general-purpose LLM. This service will be the backend for agents in our RAG application.
Initializing this service involves loading the model into vRAM and preparing the generator_model object. Given access to our cloud_volume, we can load model files from it directly, having already downloaded them using download_hf_repo from above.
@cc.service(executor=gpu_A100x1, name="Generator Service", volume=cloud_volume)
def generator_service(model_path: str):
"""Initialize the generator service.
Args:
model_path: Storage path of the generator model.
"""
from vllm import LLM
# Load the model.
generator_model = LLM(model_path)
return {
"generator_model": generator_model,
"tokenizer": generator_model.get_tokenizer(),
}
This service's /generate endpoint will then be called to generate responses. This endpoint can:
- Handle a single prompt or a batch of prompts
- Accept arbitrary sampling parameters for the
generator_modelinstance. - Apply the model's native chat template.
- Enforce an output with a specific JSON-parsable format.
@generator_service.endpoint("/generate")
def generate(
generator_model=None,
tokenizer=None,
*,
user_prompt: Optional[str] = None,
system_prompt: Optional[str] = None,
sampling_kwargs: Optional[Dict[str, Any]] = None,
schema: Optional[Dict[str, Any]] = None,
):
"""Generate text based on the user and system prompts.
Kwargs:
user_prompt: User prompt for the prompt template.
system_prompt: System prompt for the prompt template.
sampling_kwargs: Dict of kwargs passed to `vllm.SamplingParams`
schema: Optional JSON schema specifying the response format.
"""
from vllm import SamplingParams
from lmformatenforcer.integrations.vllm import (
build_vllm_logits_processor,
build_vllm_token_enforcer_tokenizer_data
)
from lmformatenforcer import JsonSchemaParser
if user_prompt is None or system_prompt is None:
return "Please provide both user and system prompts."
# Handle batched inputs.
prompts = [user_prompt] if isinstance(user_prompt, str) else user_prompt
system_prompts = [system_prompt] * len(prompts) if isinstance(system_prompt, str) else system_prompt
if not len(prompts) == len(system_prompts):
return "ERROR: The number of user and system prompts must match."
# Configure sampling parameters.
sampling_kwargs = sampling_kwargs or {"temperature": 0.8, "top_p": 0.9}
sampling_params = SamplingParams(**sampling_kwargs)
# Use the schema to enforce a specific format.
if schema:
tokenizer_data = build_vllm_token_enforcer_tokenizer_data(generator_model)
json_parser = JsonSchemaParser(schema)
logits_processor = build_vllm_logits_processor(tokenizer_data, json_parser)
sampling_params.logits_processors = [logits_processor]
# Prepare input prompts.
inputs = []
for system_prompt_, prompt_ in zip(system_prompts, prompts):
input_ids = tokenizer.apply_chat_template(
[
{"role": "system", "content": system_prompt_},
{"role": "user", "content": prompt_},
],
add_generation_prompt=True,
)
inputs.append(tokenizer.decode(input_ids))
# Generate text.
outputs = generator_model.generate(inputs, sampling_params=sampling_params)
outputs = [output.outputs[0].text for output in outputs]
return outputs if len(outputs) > 1 else outputs[0]
Workflow definition
The workflow function below (a.k.a. "the lattice") orchestrates the tasks and services we've defined above. Its purpose is to create the infrastructure for our RAG application.
As always, interdependencies are resolved automatically from the function definition, so the three calls to download_hf_repo, for example, will run concurrently. Once the generator model's repo is downloaded, the generator service will start initializing. The embedding service, on the other hand, will initialize after our Snowflake database table is populated.
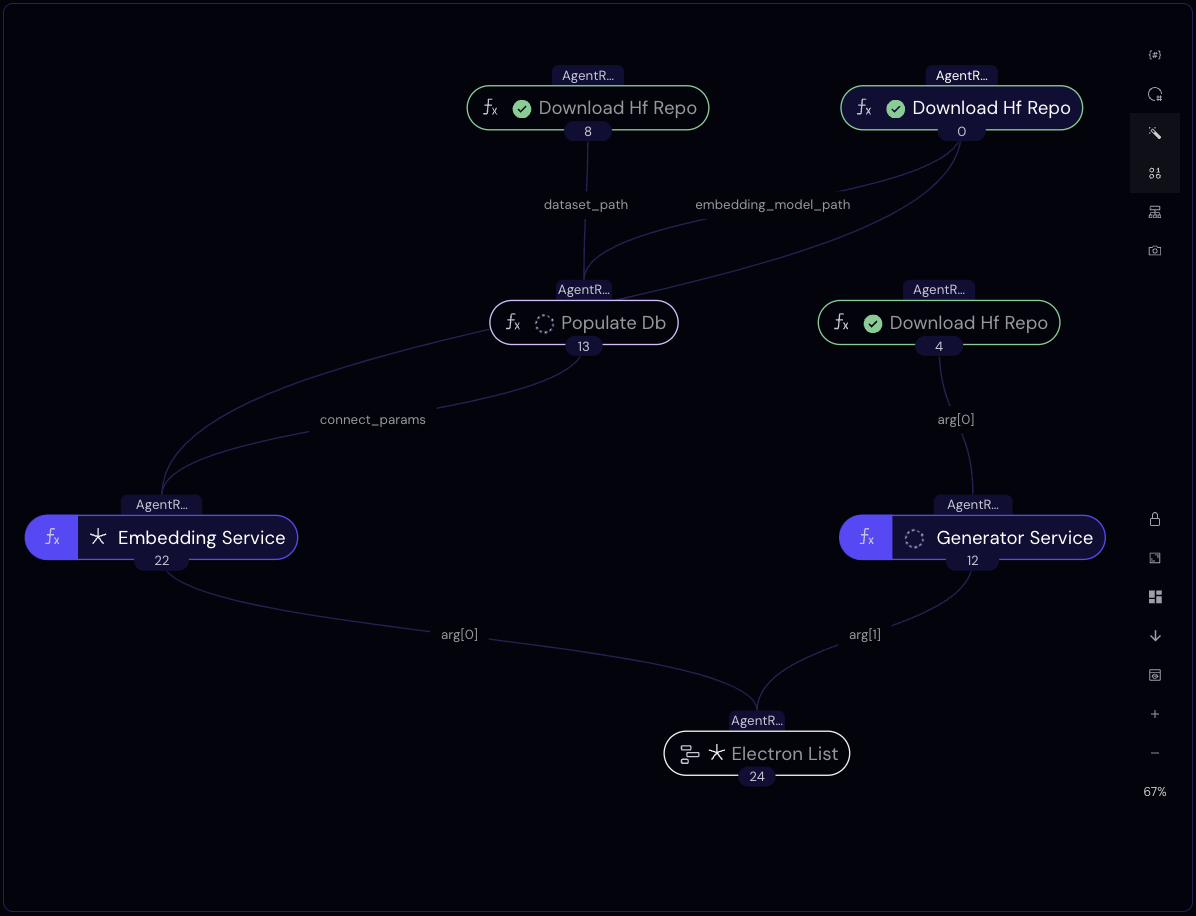
The workflow returns when both services are active.
lattice_ex = cc.CloudExecutor(
env=ENV_NAME,
num_cpus=2,
memory="16GB",
time_limit="45 minutes",
)
@ct.lattice(executor=lattice_ex, workflow_executor=lattice_ex)
def agent_rag_workflow(
embed_model_id: str,
gen_model_id: str,
dataset_id: str,
warehouse_name: str,
database_name: str,
dataset_split: str,
dataset_embed_feature: str,
dataset_name: str = "",
schema_name: str = "rag",
table_name: str = "embeddings",
overwrite_stored: bool = False,
embed_size: int = 4096,
min_items: int = 1000,
):
# Download models and dataset in parallel.
gen_model_path = download_hf_repo(
repo_id=gen_model_id, overwrite=overwrite_stored, repo_type="model"
)
generator_llm_client = generator_service(gen_model_path)
embed_model_path = download_hf_repo(
repo_id=embed_model_id, overwrite=overwrite_stored, repo_type="model"
)
dataset_path = download_hf_repo(
repo_id=dataset_id, overwrite=overwrite_stored, repo_type="dataset"
)
# Populate the database with embeddings.
connect_params = populate_db(
embedding_model_path=embed_model_path,
dataset_path=dataset_path,
dataset_name=dataset_name,
dataset_split=dataset_split,
dataset_embed_feature=dataset_embed_feature,
warehouse_name=warehouse_name,
database_name=database_name,
schema_name=schema_name,
table_name=table_name,
embed_size=embed_size,
min_items=min_items,
)
# Initialize two LLM backend services.
embedding_llm_client = embedding_service(
model_path=embed_model_path,
connect_params=connect_params,
)
return embedding_llm_client, generator_llm_client
dispatch_id = cc.dispatch(agent_rag_workflow, volume=cloud_volume)(
embed_model_id="intfloat/e5-mistral-7b-instruct",
gen_model_id="meta-llama/Meta-Llama-3-8B-Instruct",
dataset_id="rag-datasets/rag-mini-wikipedia",
dataset_name="text-corpus",
dataset_split="passages",
dataset_embed_feature="passage",
warehouse_name="rag_mini_wiki",
database_name="mini_wiki_corpus",
)
print("Workflow dispatch ID: ", dispatch_id)
Workflow dispatch ID: 54952943-b9cd-491b-bd14-b4271d7f896c
Workflow submission is asynchronous, but we can make a blocking request for the result as follows.
res = cc.get_result(dispatch_id, wait=True)
res.result.load()
Once the result is available, we unpack it to obtain clients for our two deployed services.
embed_client, gen_client = res.result.value
print(embed_client)
print(gen_client)
╭──────────────────────────────── Deployment Information ────────────────────────────────╮
│ Name Embedding Service │
│ Description Initialize the embedding service and (optionally) embed │
│ a dataset as vectors in a Snowflake DB. │
│ │
│ Args: │
│ model_path: Storage path of the embedding model. │
│ connect_params: Parameters to connect to the Snowflake DB. │
│ │
│ Function ID 66999d79880fcfb5301ee0e5 │
│ Address https://fn-c.prod.covalent.xyz/66999d79880fcfb5301ee0e5 │
│ Status ACTIVE │
│ Auth Enabled Yes │
╰────────────────────────────────────────────────────────────────────────────────────────╯
╭────────────────────────────────────── Endpoints ───────────────────────────────────────╮
│ Route POST /retrieve │
│ Streaming No │
│ Description Retrieve the most similar text from the Snowflake DB. │
│ │
│ Kwargs: │
│ text: Input text to find the most similar text. │
│ │
│ Returns: │
│ The most similar text from the Snowflake DB. │
│ │
╰────────────────────────────────────────────────────────────────────────────────────────╯
Authorization token:
qc3Fw3e5OryEn9heUhJZa93Ds8DDdDIiMps3DkPFW1dDTw-0ywdAATe4_k5wTNBg54U0Xg5OZo30khKcSJ9zbQ
╭──────────────────────────────── Deployment Information ────────────────────────────────╮
│ Name Generator Service │
│ Description Initialize the generator service. │
│ │
│ Args: │
│ model_path: Storage path of the generator model. │
│ │
│ Function ID 66999b12880fcfb5301ee0e2 │
│ Address https://fn-c.prod.covalent.xyz/66999b12880fcfb5301ee0e2 │
│ Status ACTIVE │
│ Auth Enabled Yes │
╰────────────────────────────────────────────────────────────────────────────────────────╯
╭────────────────────────────────────── Endpoints ───────────────────────────────────────╮
│ Route POST /generate │
│ Streaming No │
│ Description Generate text based on the user and system prompts. │
│ │
│ Kwargs: │
│ user_prompt: User prompt for the prompt template. │
│ system_prompt: System prompt for the prompt template. │
│ sampling_kwargs: Dict of kwargs passed to `vllm.SamplingParams` │
│ │
╰────────────────────────────────────────────────────────────────────────────────────────╯
Authorization token:
6G1FRNUgrdGptsbsipNyLR1Q1wDc_i5MEHu1d0OBath6qHkYJ_lXb9EvPfia6CFSw1Kt-9Q5x_J8QY3tZfaing
Building the RAG application
Having populated our database, deployed our embedding model, and deployed our generator LLM, we can build a RAG application relatively easily using the services that we've created.
Agents
To start, let's define an Agent class that wraps an LLM client.
from typing import Type
from pydantic import BaseModel
class Agent:
"""A simple agent that generates responses based on a role description."""
def __init__(
self,
input_template: str,
role: str,
llm_client: Deployment,
schema: Type[BaseModel] = None,
max_tokens: int = 50,
):
"""Initializes an Agent with a role and a LLM client.
Args:
input_template: A template for the input prompt.
role: A text description of the agent's role.
llm_client: An LLM client for generating responses.
schema: Optional response schema. Defaults to None.
max_tokens: Maximum number of tokens to generate. Defaults to 50.
"""
self.input_template = input_template
self.role = role
self.llm_client = llm_client
self.schema = schema
self.max_tokens = max_tokens
def system_prompt(self) -> str:
"""Returns a system prompt based on the agent's role."""
return (
"You are an AI assistant whose responses correspond "
f"to the following role description:\n{self.role}\n"
"Adhere to the role as closely as possible!"
)
def generate(self, **prompt_kwargs) -> Union[str, List[str]]:
"""Queries the LLM with a role-specific prompt."""
prompt_ = self.input_template.format(**prompt_kwargs)
sampling_kwargs = {"max_tokens": self.max_tokens}
if self.schema:
response = self.llm_client.generate(
user_prompt=prompt_,
system_prompt=self.system_prompt(),
sampling_kwargs=sampling_kwargs,
schema=self.schema.model_json_schema()
)
try:
return json.loads(response)
except json.JSONDecodeError as e:
raise ValueError(f"Error decoding response: {response}") from e
return self.llm_client.generate(
user_prompt=prompt_,
system_prompt=self.system_prompt(),
sampling_kwargs=sampling_kwargs,
)
Using the Agent class, we instantiate three agents, each with a unique role. All of these agents are conceived as wrappers on top of our generation service, which uses Llama 3.
# class GuessesList(BaseModel):
# guesses: List[str]
initial_agent = Agent( # paraphrases the initial question
input_template="Question: {question}\n",
role=(
"Given a question, generate one or more equivalent questions. "
"Generate only questions that are semantically equivalent to the original. "
"Do not answer or alter the original question.\n"
"Respond with nothing except the list of questions."
),
llm_client=gen_client,
max_tokens=100,
# schema=GuessesList, # Optional schema for response validation.
# Effectively enforces a JSON-parsable response,
# but is 2-3x slower than normal generation.
)
context_augmented_agent = Agent( # receives DB context and generates an answer
input_template="Context:\n{retrieved}\n\nQuestion:\n{question}\n",
role=(
"Generate an answer to the user's query while "
"considering the provided context. "
"Generate concise answers without additional information."
),
llm_client=gen_client,
max_tokens=20,
)
answer_evaluation_agent = Agent( # evaluates the correctness of a guessed answer
input_template=(
"Question:\n{question}\n\nGuess:\n{guess}\n\nCorrect answer:\n{answer}\n\n"
),
role=(
"You serve as a marker who, given the correct answer "
"to a question, evaluates a guessed answer to "
"mark it as either 'correct' or 'incorrect'. Guessed answers "
"are considered 'correct' if they imply the given correct answer.\n"
"You respond ONLY with either 'correct' or 'incorrect'!"
),
llm_client=gen_client,
max_tokens=10,
)
Multi-agent pipeline
The function below calls the agents in sequence, including with a query to the embedding service to retrieve relevant information. Note that the arguments use_initial_agent and use_database allow us to disable question paraphrasing and/or RAG. We'll use these arguments to compare the performance of our pipeline with and without those components.
def evaluate_with_agents(
question: str,
correct_answer: str,
use_initial_agent: bool = True,
use_database: bool = True,
verbose: bool = False,
) -> bool:
text = question
if use_initial_agent:
# Step 1: Generate equivalent questions.
equivalent_questions = initial_agent.generate(question=question)
if verbose:
print(">> Initial Agent:")
print(equivalent_questions)
print()
text = equivalent_questions
retrieved_ = ""
if use_database:
# Step 2: Retrieve relevant information.
retrieved = embed_client.retrieve(text=text)
try:
retrieved_ = "\n".join(json.loads(r)["passage"] for r in retrieved)
except json.JSONDecodeError:
retrieved_ = "\n".join(retrieved)
if verbose:
print(">> Retrieved:")
print(retrieved_)
print()
# Step 3: Generate an answer using retrieved context.
guess = context_augmented_agent.generate(
question=question, retrieved=retrieved_
)
if verbose:
print(">> LLM's guess:")
print(guess)
print()
# Step 4: Evaluate the generated answer.
correctness = answer_evaluation_agent.generate(
question=question, guess=guess, answer=correct_answer
)
if verbose:
print(">> Correctness check:")
print(correctness)
print()
print(f"[Known answer: {correct_answer}]")
return correctness in ["correct", "Correct"]
Evaluating a single example
question = "What name was given by Aristotle for the hardened shield like forewings?" # [of beetles]
known_answer = "Coleoptera"
Result without initial agent, nor RAG
evaluate_with_agents(
question,
known_answer,
use_initial_agent=False, # No initial agent to paraphrase
use_database=False, # No RAG
verbose=True,
)
>> LLM's guess:
Athenae.
>> Correctness check:
Incorrect
[Known answer: Coleoptera]
Result with complete pipeline
Let's try again with the complete pipeline.
# Complete multi-agent RAG pipeline.
evaluate_with_agents(
question,
known_answer,
use_initial_agent=True, # Use initial agent
use_database=True, # Use RAG
verbose=True,
)
>> Initial Agent:
Here are some equivalent questions:
* What term did Aristotle use to describe the hardened shield-like forewings?
* Aristotle called the hardened shield-like forewings by what name?
* What is the name coined by Aristotle for the shield-like forewings?
* What was the name assigned by Aristotle to the hardened forewings?
* What is the appellation given by Aristotle for the shield-like wing material?
>> Retrieved:
{"passage": "The name "Coleoptera" was given by Aristotle for the hardened shield like forewings (coleo = shield + ptera = wing). ", "id": 2379}
{"passage": "A cockchafer with its elytra raised, exposing the membranous flight wings, where the veins are visible", "id": 2380}
{"passage": "Acilius sulcatus, a diving beetle showing hind legs adapted for life in water", "id": 2389}
>> LLM's guess:
"Coleoptera" was given by Aristotle for the hardened shield-like forewings.
>> Correctness check:
correct
[Known answer: Coleoptera]
Evaluating many examples
Let's formalize the above to automate the evaluation process. Because the sample dataset is fairly small (12 MB), we can download it locally and run the evaluation loop using our agents, which in turn use the Python clients for our services.
pip install -q huggingface-hub datasets tqdm
huggingface-cli download rag-datasets/rag-mini-wikipedia --repo-type dataset --local-dir ./rag-mini-wikipedia --quiet
from datasets import load_dataset_builder
from tqdm import tqdm
builder = load_dataset_builder("./rag-mini-wikipedia", "question-answer")
builder.download_and_prepare()
data = builder.as_dataset(split="test")
data = data.select(range(100))
Result without initial agent, nor RAG
For comparison, we again run this without the initial agent and RAG.
total = 0
correct_count = 0
for item in tqdm(data):
question = item["question"]
correct_answer = item["answer"]
result = evaluate_with_agents(
question=question,
correct_answer=correct_answer,
use_initial_agent=False, # No initial agent
use_database=False, # No RAG
)
total += 1
correct_count += result
print(f"Score: {correct_count}/{total}")
100%|██████████| 100/100 [03:46<00:00, 2.26s/it]
Score: 56/100
Result with complete pipeline
Finally, with all the components enabled, we observe an improvement in the correctness of responses.
total = 0
correct_count = 0
for item in tqdm(data):
question = item["question"]
correct_answer = item["answer"]
result = evaluate_with_agents(
question=question,
correct_answer=correct_answer
)
total += 1
correct_count += result
print(f"Score: {correct_count}/{total}")
100%|██████████| 100/100 [22:25<00:00, 13.46s/it]
Score: 65/100
Cleaning up
Run the following to tear down the deployed services (before their executor's time_limit).
embed_client.teardown()
gen_client.teardown()
Conclusion
Using Covalent cloud, we created a workflow that (1) downloads model and data files to cloud storage, (2) runs an embedding model to populate a Snowflake database, and (3) deploys two services: one for querying embeddings and another for generation.
Using Python clients for these services, we built a RAG application that answers context-specific questions, then evaluated its performance using a sample dataset and noted a modest improvement in the correctness of responses. Curating the dataset, ensuring failure-free ingestion, and various other enhancements are likely to yield further improvements. We encourage you to tinker with this example, trying different models and/or datasets!